🧬 Dataset: BioProBench
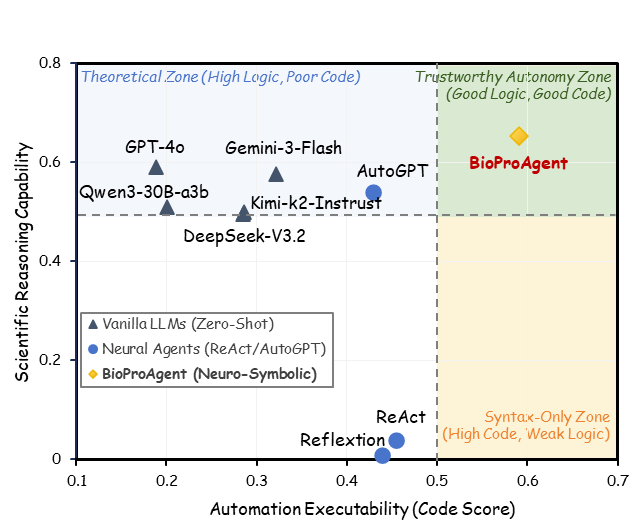
We present BioProBench, the first large-scale resource dedicated to procedural reasoning in biological experimental protocols, containing a BioProCorpus of nearly 27,000 protocols and over 550,000 structured instances, covering diverse subfields of biology.
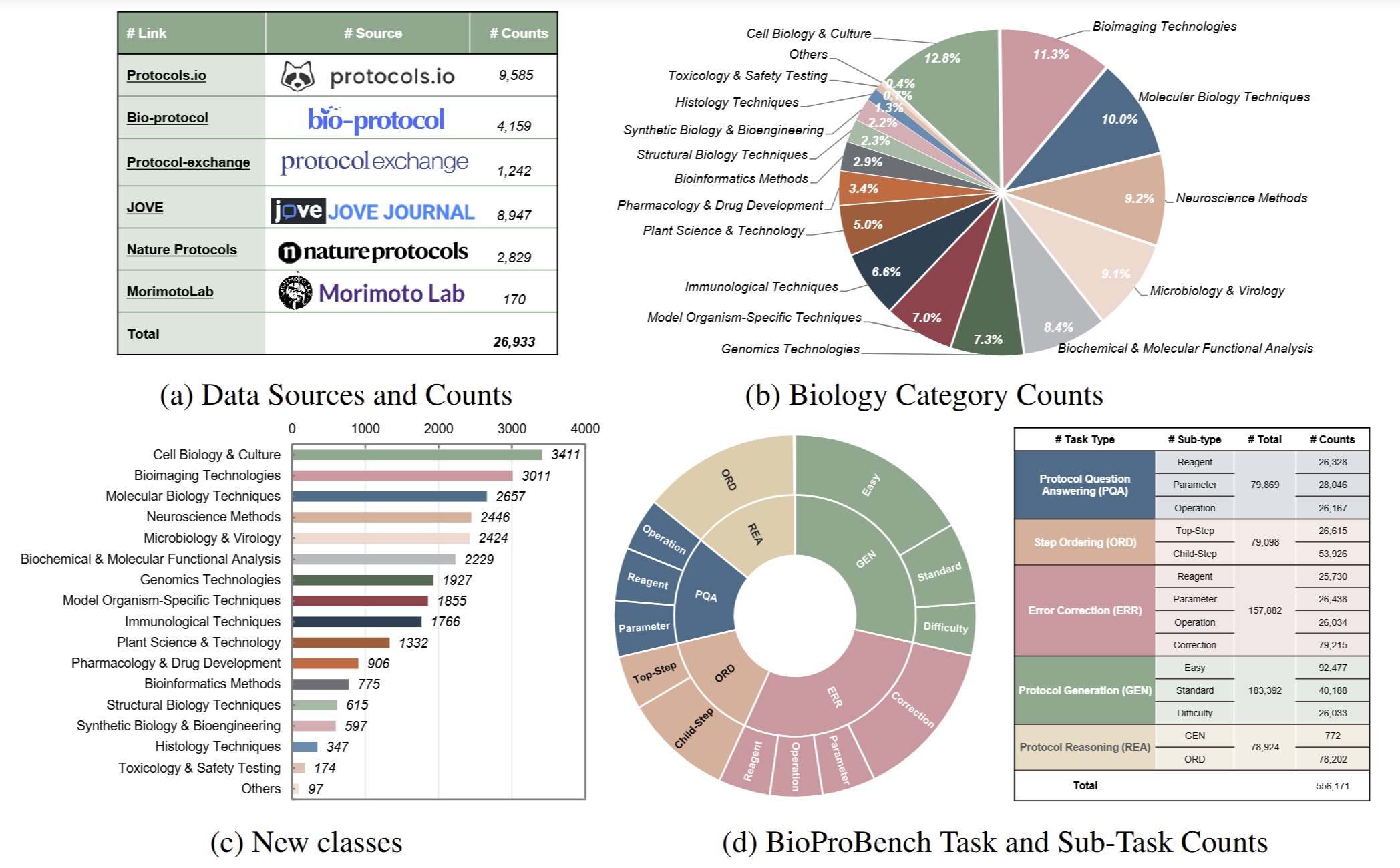
Overview of BioProBench. (a) A foundational corpus of 27,000 professionally authored protocols; (b) A structured dataset of over 550,000 instances derived from this BioProCorpus, which is partitioned into a training set to facilitate model fine-tuning and a held-out test set; and (c) A rigorous benchmark with novel, domain-specific metrics to evaluate procedural understanding, such as keyword-based content metrics and embedding-based structural metrics, to accurately quantify procedural knowledge.